Introduction
Myelin water fraction (MWF) determined by fitting a multi-exponential T2 decay curve was pioneered at UBC in the early 90s by Alex MacKay and Ken Whittall at UBC1. Since then the UBC MRI Research Centre has continued to improve, validate, and apply this technique. See the bottom of the page for the newest/fastest analysis processing of MWF data.
New software: DECAES for rapid myelin water and luminal water analysis
Recently Jon Doucette under the supervision of Dr. Alexander Rauscher (lab group page) has improved the speed of the MWI processing.
We are pleased to present DECAES, a free software tool written in Julia. It computes myelin water and luminal water maps in under two minutes. It was recently published5 and the software can be found on Jon Doucette’s Github page.
Please note: the above github page is separate from the github MWI page accessed through the submission form.
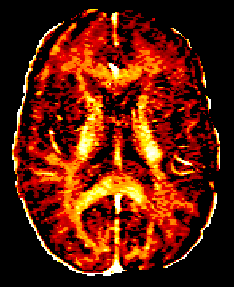
How to do Myelin Water Imaging (MWI)
Interested in running MWI at your site?
Manufacturer specific acquisition files and analysis code in MATLAB is available on the project page hosted by github (UBC MWF Project Link).
Please note you will not be able to view this page unless you’re logged into github and have been added to the member list.
In order to access this page you will need to:
1) Register an account on github (githubreg)
2) Fill out the request form here (including your github username): MWI Request Form
After completing the MWI Request Form, you will be added to the UBC MWF github project site’s member list and will then have access to files (available for download) and information regarding running MWI, as well as access to a forum to ask questions and discuss issues with implementing this technique.
How MWI works
Water trapped between the myelin bi-layers exhibits faster T2 relaxation than the intra- and extra-cellular (IE) water. Thus, if the T2 decay curve is sampled at many echo times, the contributions of both the myelin water and IE water fractions can be separated with an NNLS fit1,2.
Implementations of MWI are based on a 3D gradient and spin echo (GraSE) sequence with parallel imaging to reduce the acquisition time to just under 8 minutes for whole cerebrum coverage3.
References
1.https://doi.org/10.1002/mrm.1910310614
2.https://doi.org/10.1016/0022-2364(89)90011-5
3.https://doi.org/10.1016/j.neuroimage.2012.06.064
